| age | outdoor_time | social_int | routine | wellbeing | location | steps_k |
|---|---|---|---|---|---|---|
| 28 | 12 | 13 | 1 | 36 | rural | 21.6 |
| 56 | 5 | 15 | 1 | 41 | rural | 12.3 |
| 25 | 19 | 11 | 1 | 35 | rural | 49.8 |
| 60 | 25 | 15 | 0 | 35 | rural | NA |
| 19 | 9 | 18 | 1 | 32 | rural | 48.1 |
| 34 | 18 | 13 | 1 | 34 | rural | 67.3 |
Simple Linear Regression
Learning Objectives
At the end of this lab, you will:
- Be able to specify a simple linear model.
- Understand what fitted values and residuals are.
- Be able to interpret the coefficients of a fitted model.
Requirements
- Be up to date with lectures from Weeks 1 & 2
- Have completed Week 1 lab exercises
Required R Packages
Remember to load all packages within a code chunk at the start of your RMarkdown file using library(). If you do not have a package and need to install, do so within the console using install.packages(" "). For further guidance on installing/updating packages, see Section C here.
For this lab, you will need to load the following package(s):
- tidyverse
- sjPlot
Lab Data
You can download the data required for this lab here or read it in via this link https://uoepsy.github.io/data/riverview.csv.
Study Overview
Research Question
Is there an overall effect of the number of social interactions on wellbeing scores?
Setup
- Create a new RMarkdown file
- Load the required package(s)
- Read the wellbeing dataset into R, assigning it to an object named
mwdata
Exercises
Data Exploration
The common first port of call for almost any statistical analysis is to explore the data, and we can do this visually and/or numerically.
| ================ Description | Marginal Distributions | Bivariate Associations | =================================================================================================================================================================+==============================================================================================================================================================================+ The distribution of each variable without reference to the values of the other variables | Describing the relationship between two numeric variables | | ||
|
Visually |
Plot each variable individually. You could use, for example, geom_density() for a density plot or geom_histogram() for a histogram to comment on and/or examine:
|
Plot associations among two variables. | | | You could use, for example, geom_point() for a scatterplot to comment on and/or examine: | | |
|
|
Numerically |
Compute and report summary statistics e.g., mean, standard deviation, median, min, max, etc. You could, for example, calculate summary statistics such as the mean ( mean()) and standard deviation (sd()), etc. within summarize()
|
Compute and report the correlation coefficient. You can use the cor() function to calculate this |
|
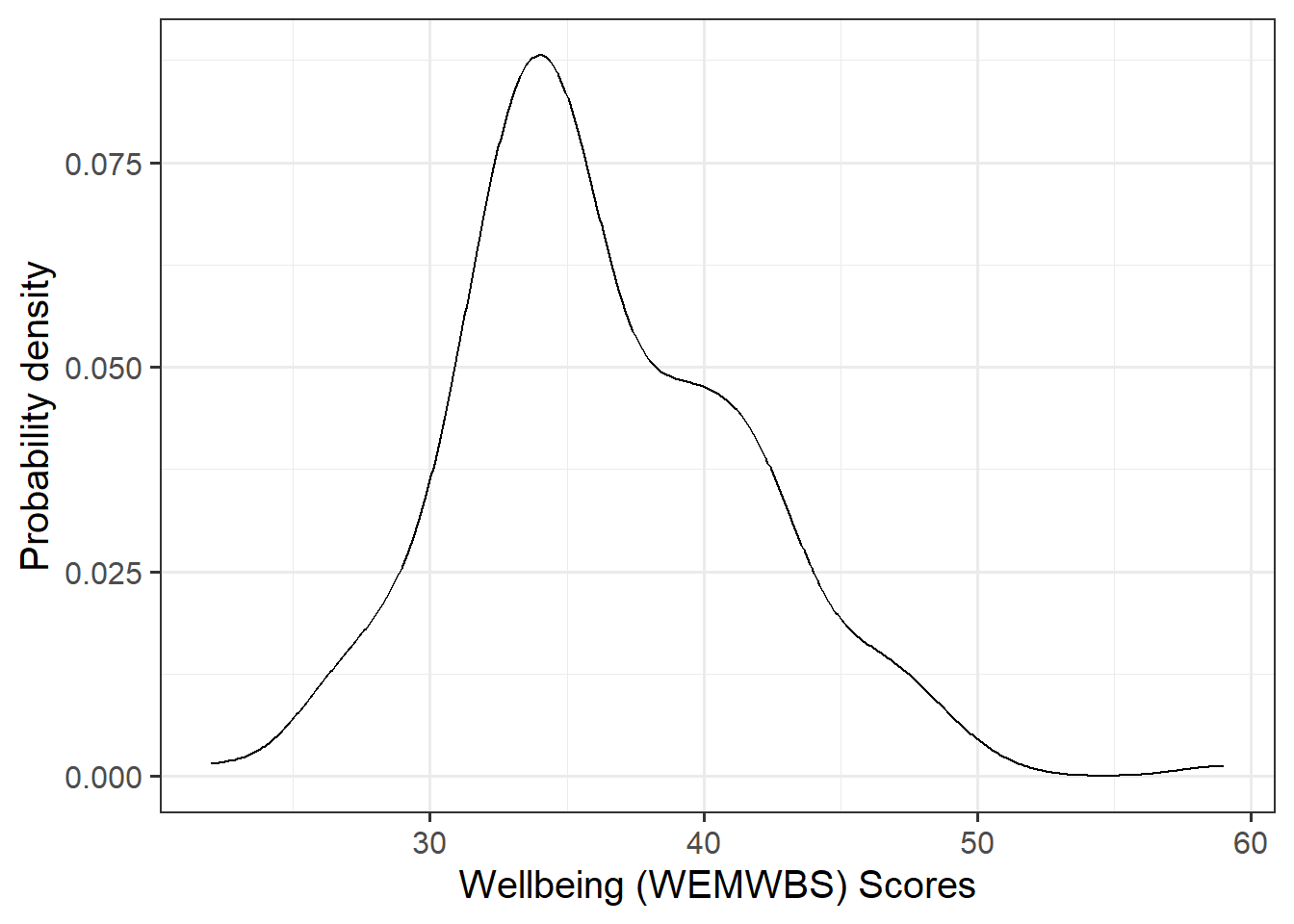
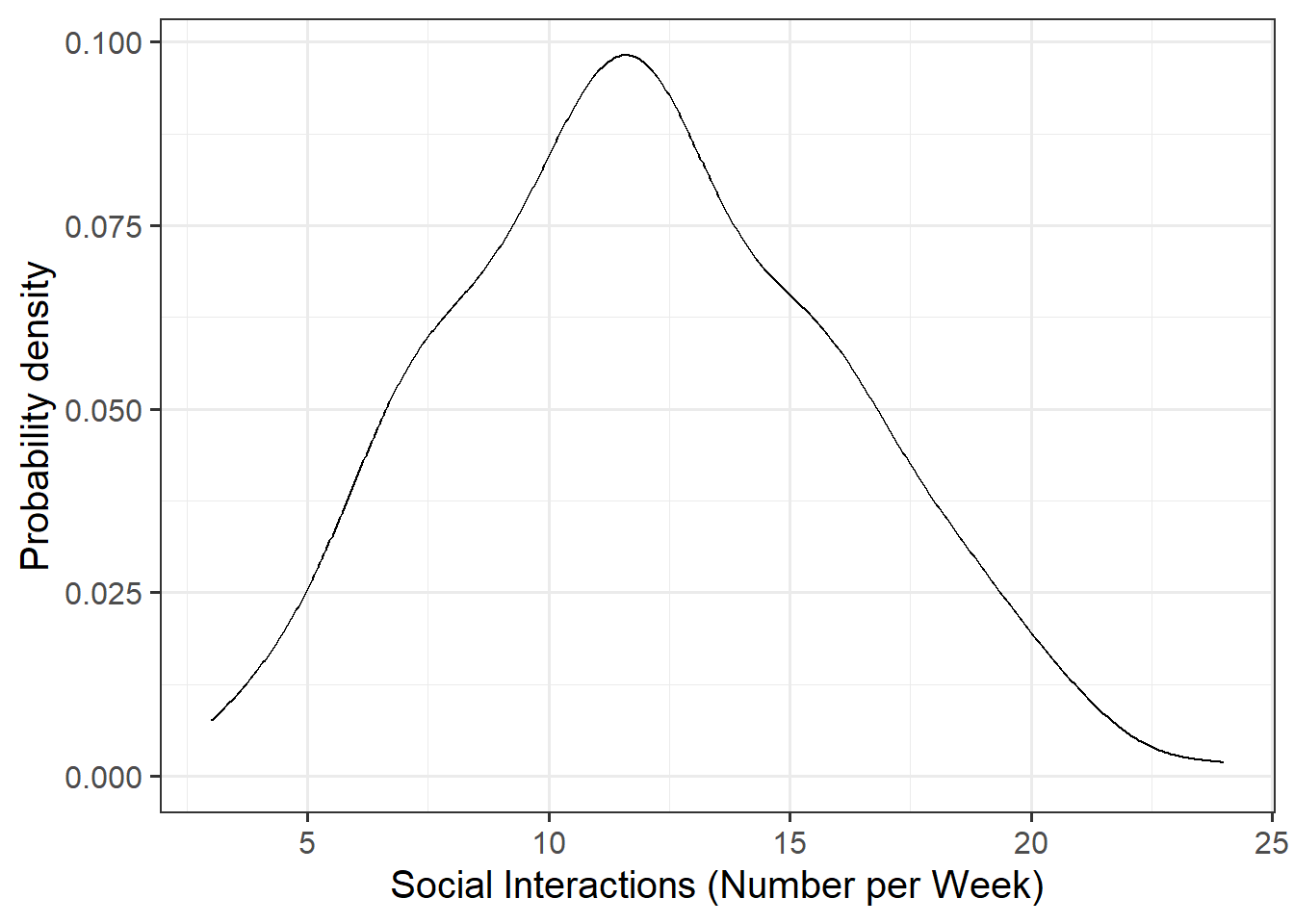
Marginal Distributions
Visualise and describe the marginal distributions of wellbeing scores and social interactions.
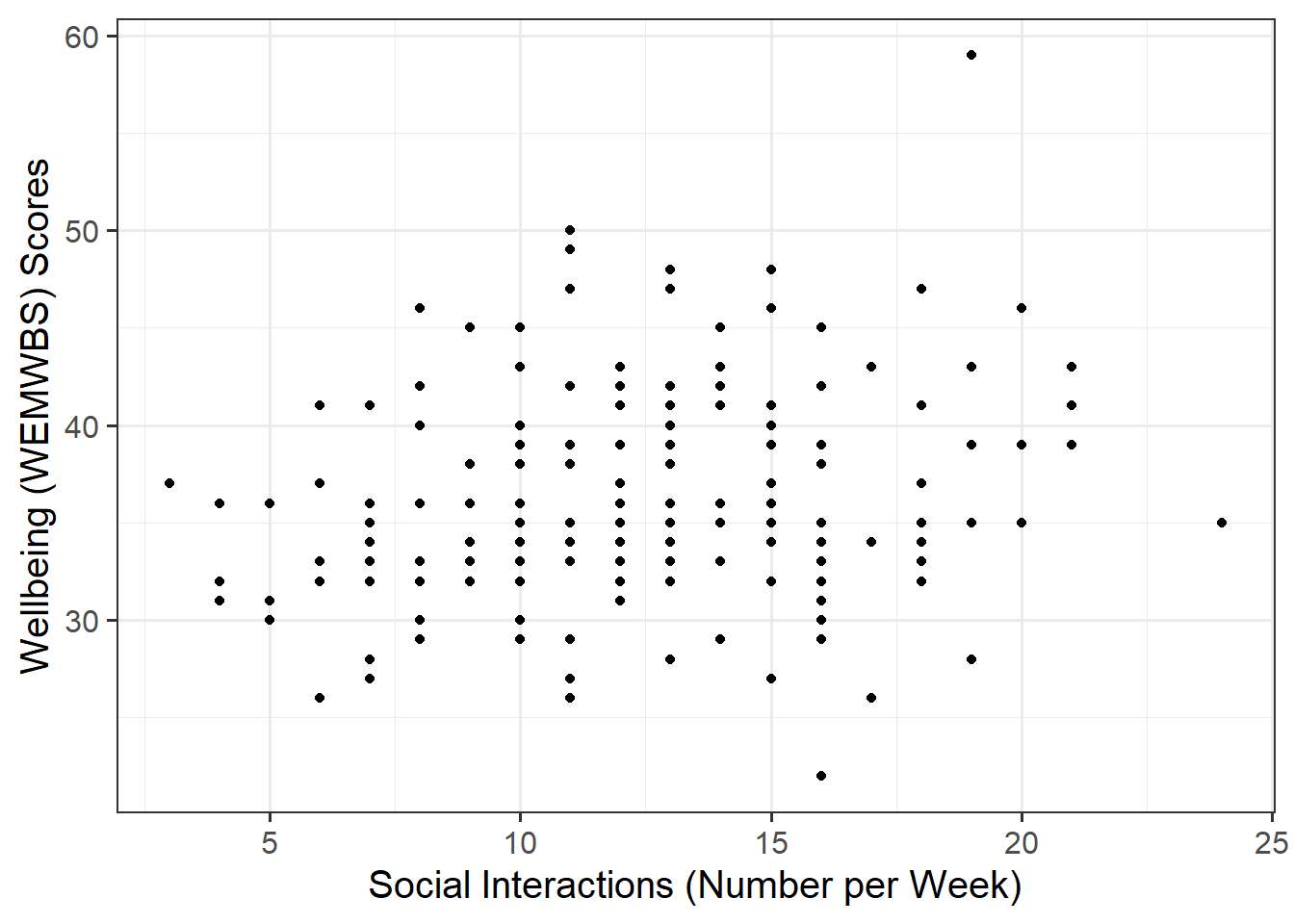
Associations among Variables
Create a scatterplot of wellbeing score and social interactions before calculating the correlation between them.
Correlation Matrix
A table showing the correlation coefficients - \(r_{(x,y)}=\frac{\mathrm{cov}(x,y)}{s_xs_y}\) - between variables. Each cell in the table shows the association between two variables. The diagonals show the correlation of a variable with itself (and are therefore always equal to 1).
Making reference to both the plot and correlation coefficient, describe the association between wellbeing and social interactions among participants in the Edinburgh & Lothians sample.
Plot
We are trying to investigate how wellbeing varies by varying numbers of weekly social interactions. Hence, wellbeing is the dependent variable (on the y-axis), and social interactions is the independent variable (on the x-axis).
Correlation
Make sure to round your numbers in-line with APA 7th edition guidelines. The round() function will come in handy here, as might this APA numbers and statistics guide!
Model Specification and Fitting
The scatterplot highlighted a linear relationship, where the data points were scattered around an underlying linear pattern with a roughly-constant spread as x varied.
Hence, we will try to fit a simple (i.e., one x variable only) linear regression model:
\[ y_i = \beta_0 + \beta_1 x_i + \epsilon_i \\ \quad \text{where} \quad \epsilon_i \sim N(0, \sigma) \text{ independently} \]
First, write the equation of the fitted line.
Next, using the lm() function, fit a simple linear model to predict wellbeing (DV) by social interactions (IV), naming the output mdl.
Lastly, update your equation of the fitted line to include the estimated coefficients.
The syntax of the lm() function is:
[model name] <- lm([response variable i.e., dependent variable] ~ [explanatory variable i.e., independent variable], data = [dataframe])
Explore the following equivalent ways to obtain the estimated regression coefficients — that is, \(\hat \beta_0\) and \(\hat \beta_1\) — from the fitted model:
mdlmdl$coefficientscoef(mdl)coefficients(mdl)summary(mdl)
Explore the following equivalent ways to obtain the estimated standard deviation of the errors — that is, \(\hat \sigma\) — from the fitted model mdl:
sigma(mdl)summary(mdl)
Interpret the estimated intercept, slope, and standard deviation of the errors in the context of the research question.
To interpret the estimated standard deviation of the errors we can use the fact that about 95% of values from a normal distribution fall within two standard deviations of the center.
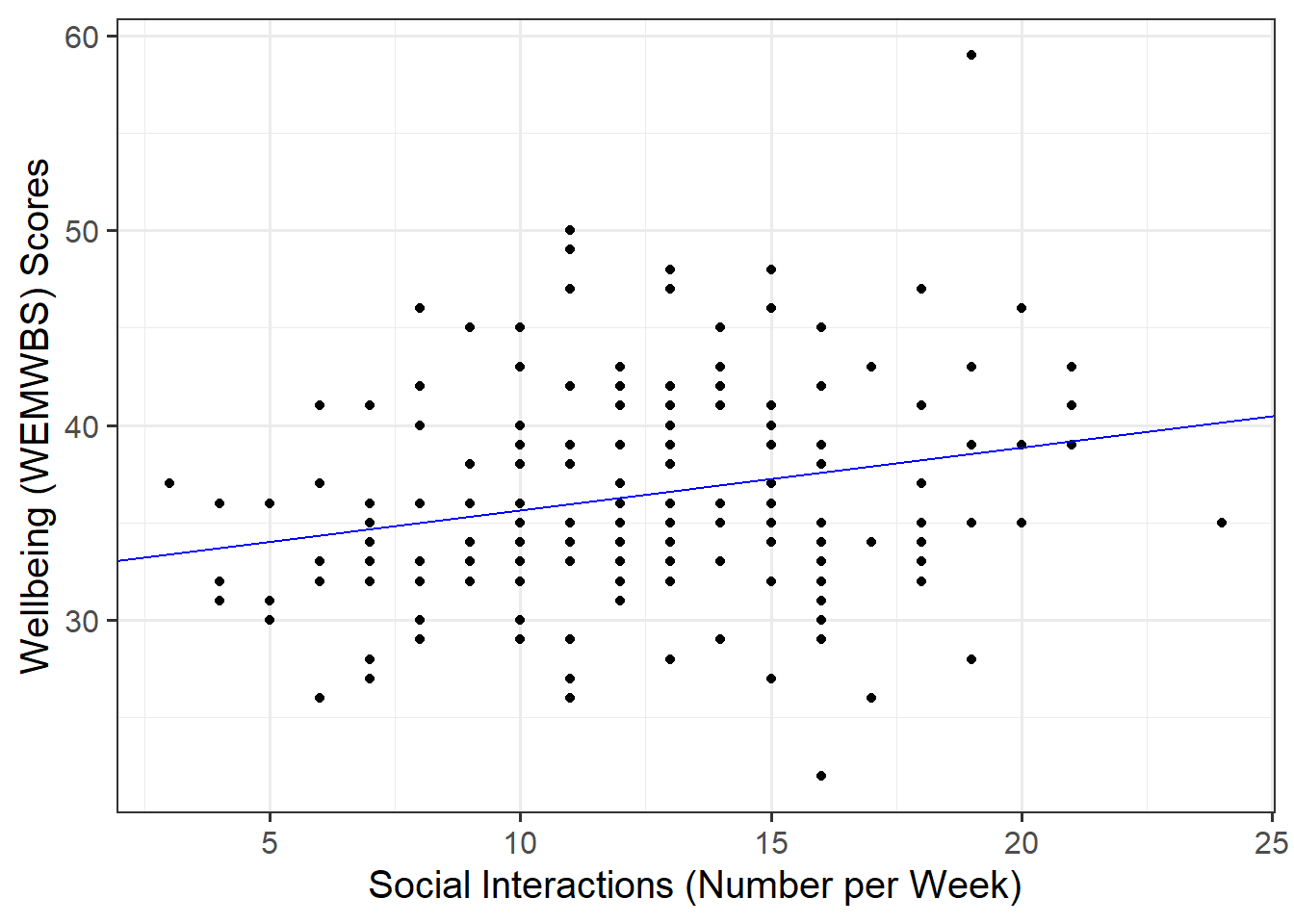
Plot the data and the fitted regression line. To do so:
- Extract the estimated regression coefficients e.g., via
betas <- coef(mdl) - Extract the first entry of
betas(i.e., the intercept) viabetas[1] - Extract the second entry of
betas(i.e., the slope) viabetas[2] - Provide the intercept and slope to the function
Extracting values
The function coef(mdl) returns a vector (a sequence of numbers all of the same type). To get the first element of the sequence you append [1], and [2] for the second.
Plotting
In your ggplot(), you will need to specify geom_abline(). This might help get you started:
geom_abline(intercept = <intercept>, slope = <slope>)
Predicted Values & Residuals
Predicted Values
Model predicted values for sample data:
We can get out the model predicted values for \(y\), the “y hats” (\(\hat y\)), for the data in the sample using various functions:
predict(<fitted model>)fitted(<fitted model>)fitted.values(<fitted model>)mdl$fitted.values
For example, this will give us the estimated wellbeing score (point on our regression line) for each observed value of social interactions for each of our 200 participants.
predict(mdl) 1 2 3 4 5 6 7 8
36.59625 37.24064 35.95186 37.24064 38.20723 36.59625 38.52943 36.27406
9 10 11 12 13 14 15 16
36.59625 37.24064 34.66308 35.62967 34.66308 36.27406 36.91845 37.88504
17 18 19 20 21 22 23 24
37.56284 37.56284 35.62967 35.30747 35.62967 38.20723 36.27406 35.30747
25 26 27 28 29 30 31 32
38.52943 38.52943 37.56284 36.27406 37.56284 35.30747 37.56284 36.91845
33 34 35 36 37 38 39 40
37.56284 34.34088 37.56284 36.27406 36.27406 36.91845 38.85162 35.30747
41 42 43 44 45 46 47 48
38.20723 36.59625 37.56284 36.27406 36.27406 35.95186 34.34088 38.85162
49 50 51 52 53 54 55 56
35.62967 37.24064 35.62967 34.98527 37.56284 37.24064 34.01869 35.95186
57 58 59 60 61 62 63 64
34.66308 37.88504 36.91845 34.66308 37.88504 35.95186 36.27406 35.95186
65 66 67 68 69 70 71 72
36.27406 36.91845 38.20723 34.01869 35.30747 35.30747 34.66308 39.17382
73 74 75 76 77 78 79 80
34.01869 35.30747 33.69649 38.52943 35.62967 37.56284 39.17382 36.91845
81 82 83 84 85 86 87 88
34.98527 35.95186 36.59625 36.27406 34.66308 35.95186 37.24064 37.88504
89 90 91 92 93 94 95 96
35.62967 37.56284 34.66308 34.66308 37.24064 35.95186 34.98527 35.62967
97 98 99 100 101 102 103 104
35.95186 34.66308 37.24064 35.95186 34.34088 34.66308 37.56284 34.98527
105 106 107 108 109 110 111 112
36.27406 37.88504 39.17382 37.24064 38.52943 35.30747 35.62967 35.95186
113 114 115 116 117 118 119 120
37.24064 36.27406 36.27406 36.59625 36.27406 35.95186 36.59625 35.95186
121 122 123 124 125 126 127 128
37.56284 36.91845 34.34088 34.66308 34.98527 35.95186 34.01869 35.62967
129 130 131 132 133 134 135 136
34.98527 35.62967 33.69649 38.20723 38.20723 35.30747 34.66308 37.56284
137 138 139 140 141 142 143 144
36.91845 35.62967 36.91845 38.52943 36.91845 35.30747 35.62967 37.88504
145 146 147 148 149 150 151 152
36.27406 34.98527 35.30747 36.91845 36.59625 36.59625 35.95186 34.34088
153 154 155 156 157 158 159 160
36.59625 36.59625 34.66308 36.91845 36.27406 35.30747 33.69649 35.62967
161 162 163 164 165 166 167 168
36.27406 37.24064 35.95186 36.59625 33.37429 36.59625 38.20723 36.27406
169 170 171 172 173 174 175 176
38.85162 34.66308 37.24064 35.62967 38.20723 36.27406 36.59625 40.14041
177 178 179 180 181 182 183 184
35.30747 34.98527 34.34088 35.95186 36.59625 34.98527 34.98527 35.62967
185 186 187 188 189 190 191 192
34.66308 34.98527 36.27406 37.88504 36.27406 35.95186 35.95186 36.59625
193 194 195 196 197 198 199 200
37.56284 36.27406 36.27406 35.95186 34.66308 35.62967 36.27406 37.24064 Model predicted values for other (unobserved) data:
To compute the model-predicted values for unobserved data (i.e., data not contained in the sample), we can use the following function:
predict(<fitted model>, newdata = <dataframe>)
For this example, we first need to remember that the model predicts wellbeing using the independent variable social_int. Hence, if we want predictions for new (unobserved) data, we first need to create a tibble with a column called social_int containing the number of weekly social interactions for which we want the prediction, and store this as a dataframe.
#Create dataframe 'newdata' containing 2, 25, and 28 weekly social interactions
newdata <- tibble(social_int = c(2, 25, 28))
newdata# A tibble: 3 × 1
social_int
<dbl>
1 2
2 25
3 28Then we take newdata and add a new column called wellbeing_hat, computed as the prediction from the fitted mdl using the newdata above:
Residuals
The residuals represent the deviations between the actual responses and the predicted responses and can be obtained either as
mdl$residualsresid(mdl)residuals(mdl)- computing them as the difference between the response (\(y_i\)) and the predicted response (\(\hat y_i\))
Use predict(mdl) to compute the fitted values and residuals. Mutate the mwdata dataframe to include the fitted values and residuals as extra columns.
Assign to the following symbols the corresponding numerical values:
- \(y_{3}\) (response variable for unit \(i = 3\) in the sample data)
- \(\hat y_{3}\) (fitted value for the third unit)
- \(\hat \epsilon_{5}\) (the residual corresponding to the 5th unit, i.e., \(y_{5} - \hat y_{5}\))
Provide key model results in a formatted table.
Use tab_model() from the sjPlot package.
You can rename your DV and IV labels by specifying dv.labels and pred.labels. To do so, specify your variable name on the left, and what you would like this to be named in the table on the right.
Describe the design of the study, and the analyses that you undertook. Interpret your results in the context of the research question and report your model results in full.
Make reference to your descriptive plots and/or statistics and regression table.
Make sure to write your results up following APA guidelines