| variable | description |
|---|---|
| Diagnosis | Diagnosis classifies the three types of individuals: 1 = Amnesic patients, 2 = Huntingtons patients, and 3 = Control group of individuals with no known neurological disorder |
| Task | Task tells us to which one of three tasks each study participant was randomly assigned to: 1 = Grammar (which consists of classifying letter sequences as either following or not following grammatical rules), 2 = Classification (which consists of classifying stimuli into certain groupings, based on previously indicated information about the groups characteristics), and 3 = Recognition (which consists of recognising particular stimuli as stimuli that have previously been presented during the task) |
| Y | Score |
Manual Contrast Interactions, Simple Effects, & Corrections
Learning Objectives
At the end of this lab, you will:
- Understand how to interpret simple effects
- Understand how to test interactions between manual contrasts
- Understand how to apply corrections for multiple comparisons
What You Need
- Be up to date with lectures
- Have completed previous lab exercises from Semester 1 Week 7, Semester 1 Week 8, Semester 1 Week 11, Semester 2 Week 1, Semester 2 Week 2, and Semester 2 Week 3.
Required R Packages
Remember to load all packages within a code chunk at the start of your RMarkdown file using library(). If you do not have a package and need to install, do so within the console using install.packages(" "). For further guidance on installing/updating packages, see Section C here.
For this lab, you will need to load the following package(s):
- tidyverse
- psych
- kableExtra
- sjPlot
- interactions
- patchwork
- emmeans
Presenting Results
All results should be presented following APA guidelines.If you need a reminder on how to hide code, format tables/plots, etc., make sure to review the rmd bootcamp.
The example write-up sections included as part of the solutions are not perfect - they instead should give you a good example of what information you should include and how to structure this. Note that you must not copy any of the write-ups included below for future reports - if you do, you will be committing plagiarism, and this type of academic misconduct is taken very seriously by the University. You can find out more here.
Lab Data
You can download the data required for this lab here or read it in via this link https://uoepsy.github.io/data/cognitive_experiment.csv
Note, you have already worked with some of this data last week - see Semester 2 Week 3 lab, but we now have a third Task condition - Classification.
Study Overview
Research Question
Are there differences in types of memory deficits for those experiencing different cognitive impairment(s)?
And we’ll use an interaction between manual contrasts to specifically ask:
Is there a difference between the effects of implicit and explicit memory tasks for amnesic patients and for Huntington’s patients?
Setup
- Create a new RMarkdown file
- Load the required package(s)
- Read the cognitive_experiment dataset into R, assigning it to an object named
cog
Exercises
Study & Analysis Plan Overview (max 15 minutes)
(You should aim to spend max five minutes on this question.)
Firstly, examine the dataset, and perform any necessary and appropriate data management steps.
Next, we will use treatment coding for both predictors:
- For
Diagnosis, the reference level will becontrol. - For
Task, the reference level will berecognition.
In your code, set these levels as the reference levels.
Data Management
- The
str()function will return the overall structure of the dataset, this can be quite handy to look at
- Convert categorical variables to factors, and if needed, provide better variable names*
- Label factors appropriately to aid with your model interpretations if required*
- Check that the dataset is complete (i.e., are there any
NAvalues?). We can check this usingis.na()
Note that all of these steps can be done in combination - the mutate() and factor() functions will likely be useful here.
A priori contrast coding schemes
- If you think you’d benefit from a refresher on contrast coding schemes, it might be best to revisit the materials from Semester 1 Block 2 (especially the dummy vs effects coding flashcard).
- If you would like an overview of coding constraints in the context of interaction models, review the categorical x categorical example > coding constraints flashcard.
Reference Levels
- Review the specifying reference levels flashcard.
*See the numeric outcomes & categorical predictors flashcard.
(You should aim to spend max ten minutes on this question.)
Provide a brief overview of the study design and data, before detailing your analysis plan to address the research question.
- Give the reader some background on the context of the study (you might be able to re-use some of the content you wrote for Semester 2 Week 3 lab here, but note that we now have an extra condition within Task)
- Outline data checks / data cleaning
- State what type of analysis you will conduct in order to address the research question
- Specify the model to be fitted to address the research question (note that you will need to specify the reference level of your categorical variables. This will be somewhat similar to last week, but with the addition of Classification in Task, our model will contain a different number of parameters)
- Specify your chosen significance (\(\alpha\)) level
- State your hypotheses
Much of the information required can be found in the Study Overview codebook.
The statistical models flashcards may also be useful to refer to. Specifically the interaction models flashcards and categorical x categorical example flashcards might be of most use.
Descriptive Statistics & Visualisations (approx 5 minutes)
Provide a table of descriptive statistics.
Interpret the descriptive statistics in the context of the study (i.e., comment on any observed differences among groups).
Review the many ways to numerically and visually explore your data by reading over the data exploration flashcards.
For examples, see flashcards on descriptives statistics tables - categorical and numeric values examples.
Make sure to comment on any observed differences among the sample means of the different conditions.
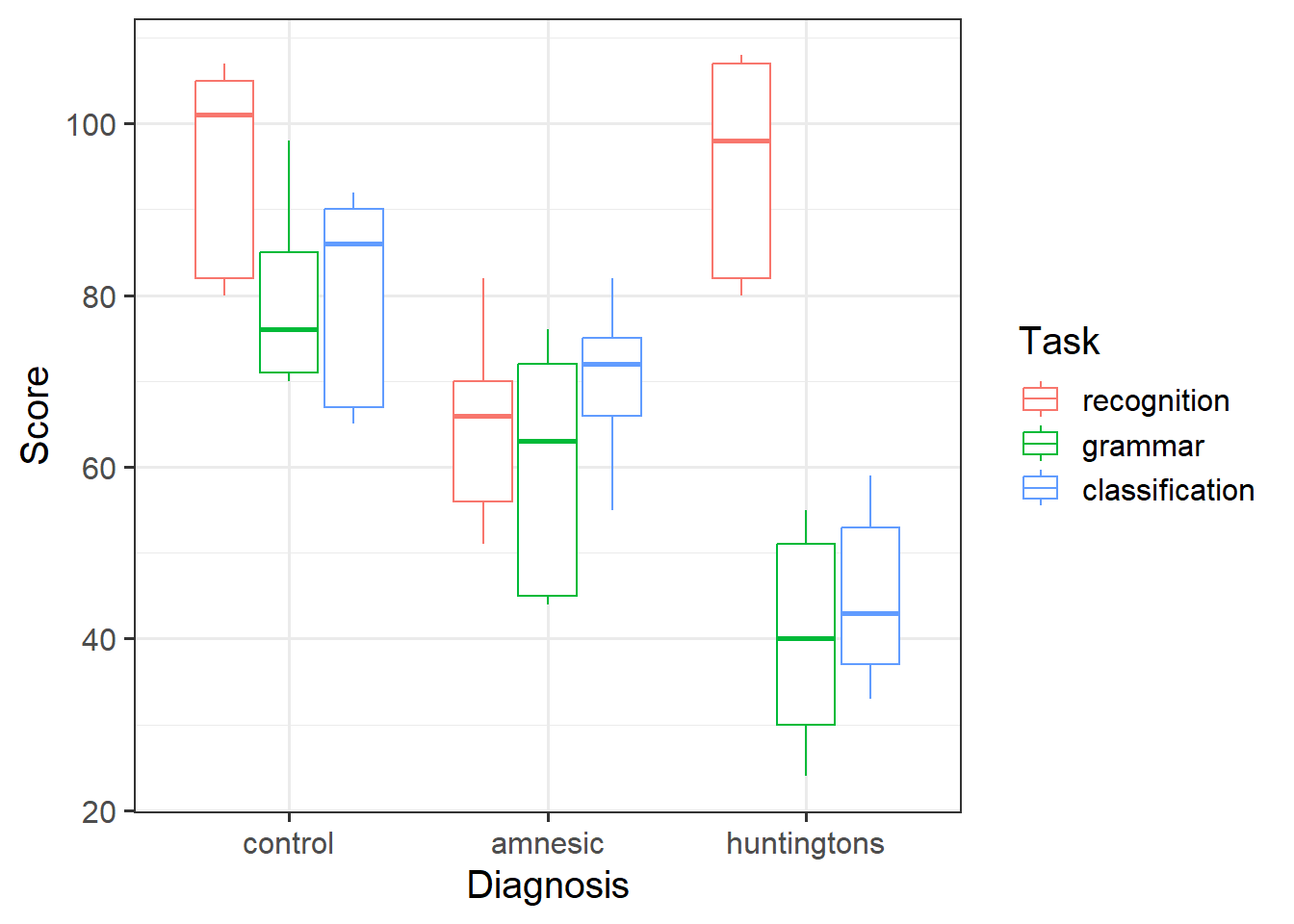
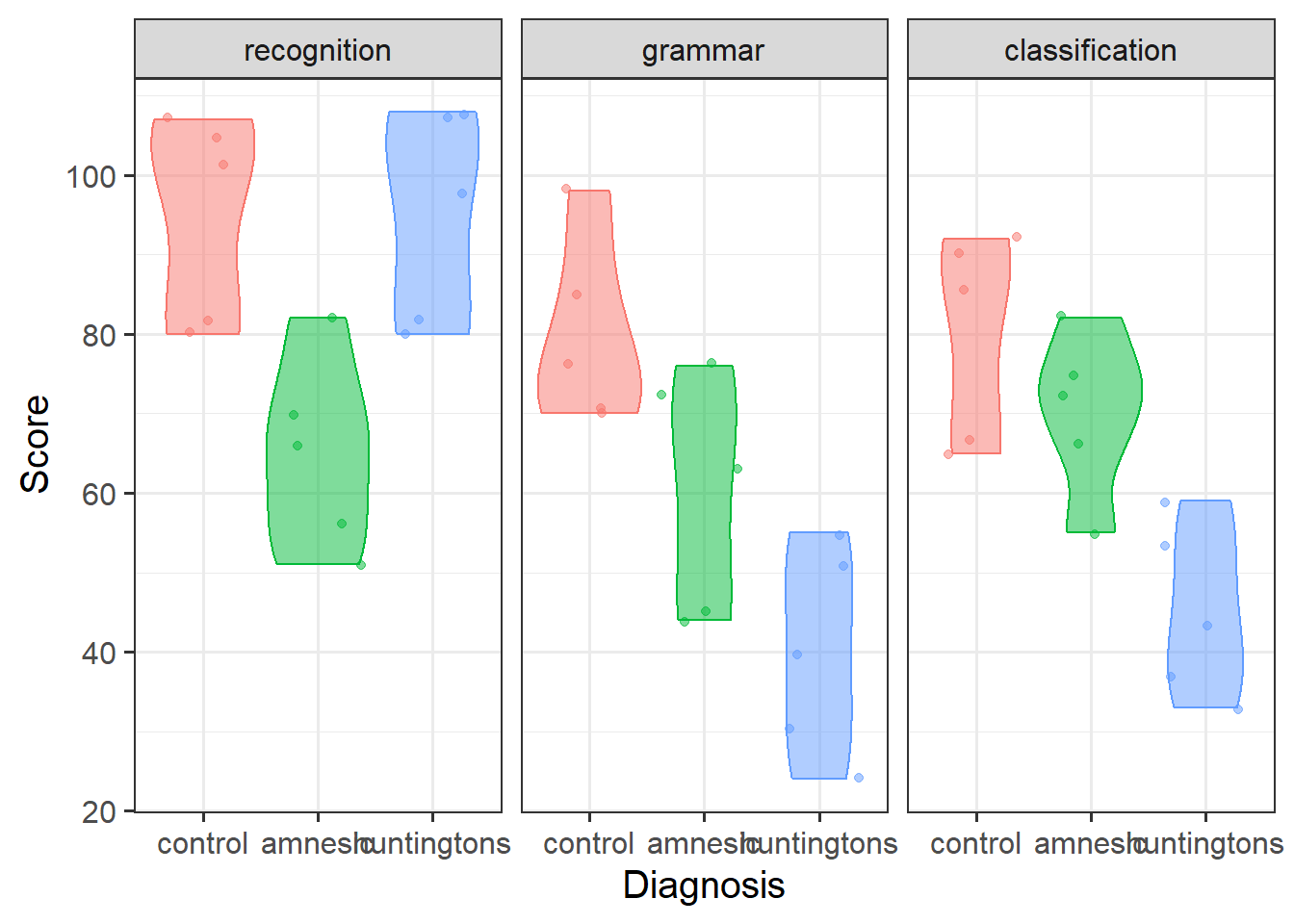
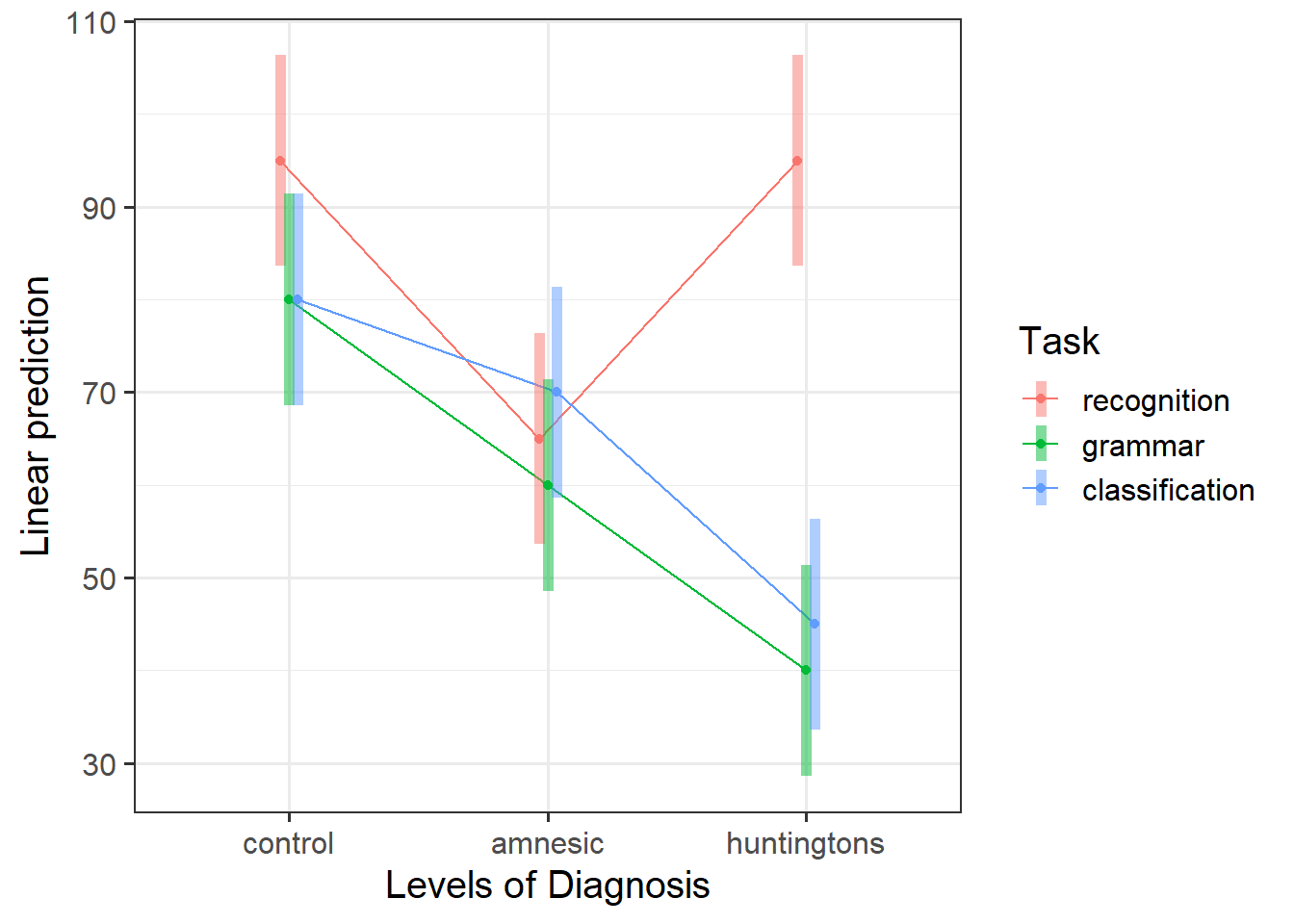
Visualise the data. As a sense check, make sure the visualisation captures the same patterns you observed in the descriptive statistics you just computed.
Review the many ways to numerically and visually explore your data by reading over the data exploration flashcards.
For examples, see flashcards on categorical x categorical example - visualise data.
Model Fitting (approx 2 min)
Fit the specified model using lm(), and assign it the name “mdl_int”.
Model Building
- We can fit interaction models using the
lm()function.
- For an overview, see the interaction models flashcards.
- For an example, review the interaction models > categorical x categorical example > model building flashcards.
Results Table (not mandatory for today)
- Use
tab_model()from the sjPlot package. For a quick guide, review the tables flashcard.
Plot Model (not mandatory for today)
- Using the
cat_plot()function from the interactions package, visualise the interaction effects from your model. - For an overview and example, review the interaction models > categorical x categorical example > model visualisation flashcards.
Interactions between manual contrasts (approx 15–20 min)
Let’s move onto to the main focus of the lab: testing differences between manually-specified differences of group means.
Your task in this question is to define the interaction term’s contrast, based on the contrasts given for the diagnostic groups and tasks.
In terms of the diagnostic groups, we want to compare the individuals with amnesia to those with Huntingtons, ignoring the control group. This corresponds to a contrast with coefficients of 0, 1, and −1, for control, amnesic, and Huntingtons, respectively.
Similarly, in terms of the tasks, we want to compare the average of the two implicit memory tasks with the explicit memory task. This corresponds to a contrast with coefficients of 0.5, 0.5, and −1 for the three tasks.
To figure out the coding values for an interaction’s contrast, we need to multiply all combinations of the manual contrast’s coding values together. This is shown schematically below:
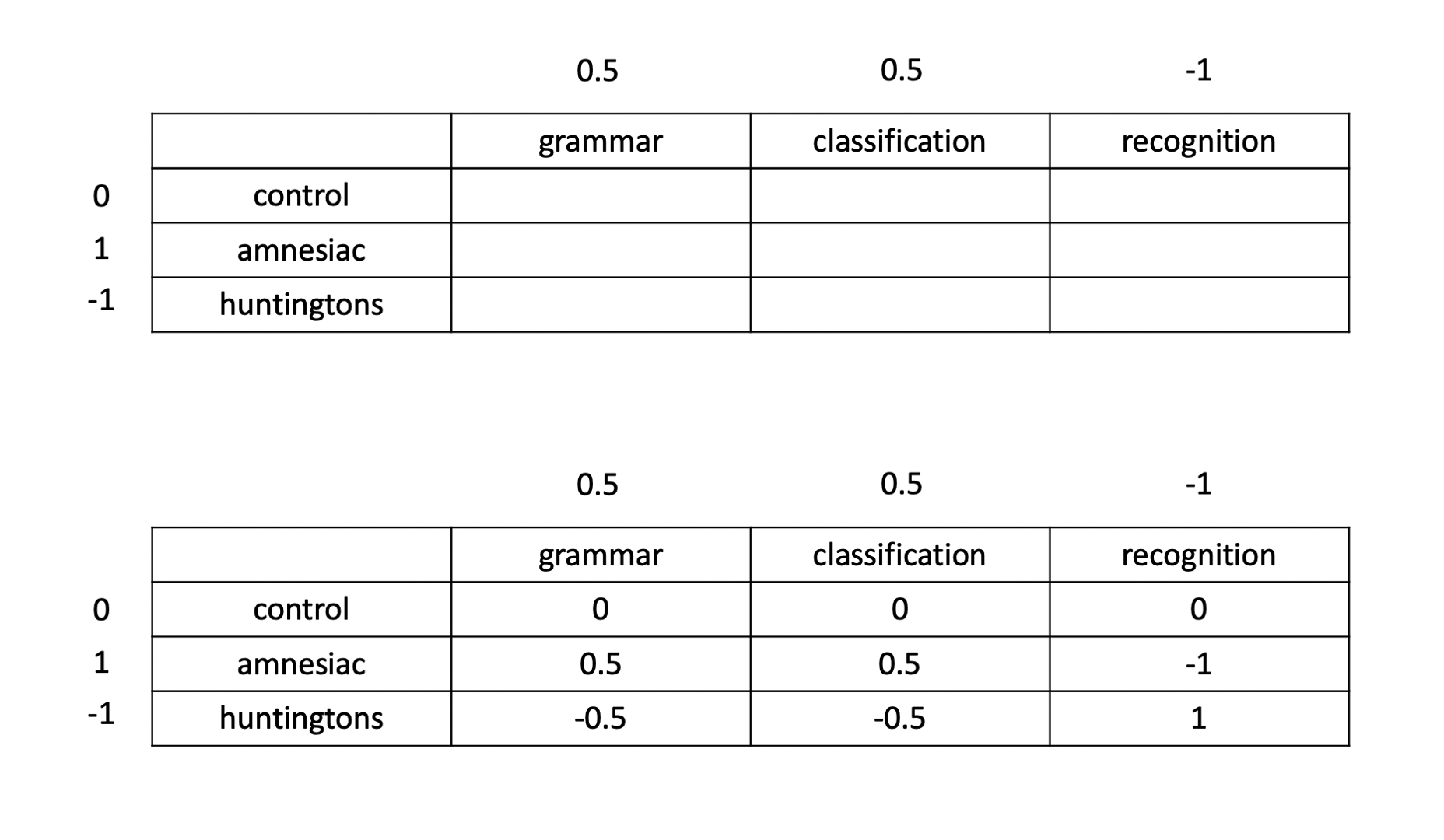
Specify the interaction term’s contrast in R. Present it in a well-formatted table that contains the same information as the one above.
For an overview and example, review the manual contrasts flashcards > interaction models.
Formally state the contrast that the researchers are interested in as testable hypotheses.
Firstly, use emmeans() to obtain the estimated means and uncertainties for all combinations of factor levels.
Next, specify the coefficients of the comparison—that is, format the contrast for the interaction term the way that emmeans expects. Run the contrast analysis, obtaining 95% confidence intervals for the interaction term.
Report the results of the contrast analysis in full.
For an overview and example, review the manual contrasts flashcards > interaction models.
Simple Effects (approx 10 min)
Examine the simple effects for Task at each level of Diagnosis; and then the simple effects for Diagnosis at each level of Task.
For an overview and example, review the interaction models > categorical x categorical example > simple effects flashcards.
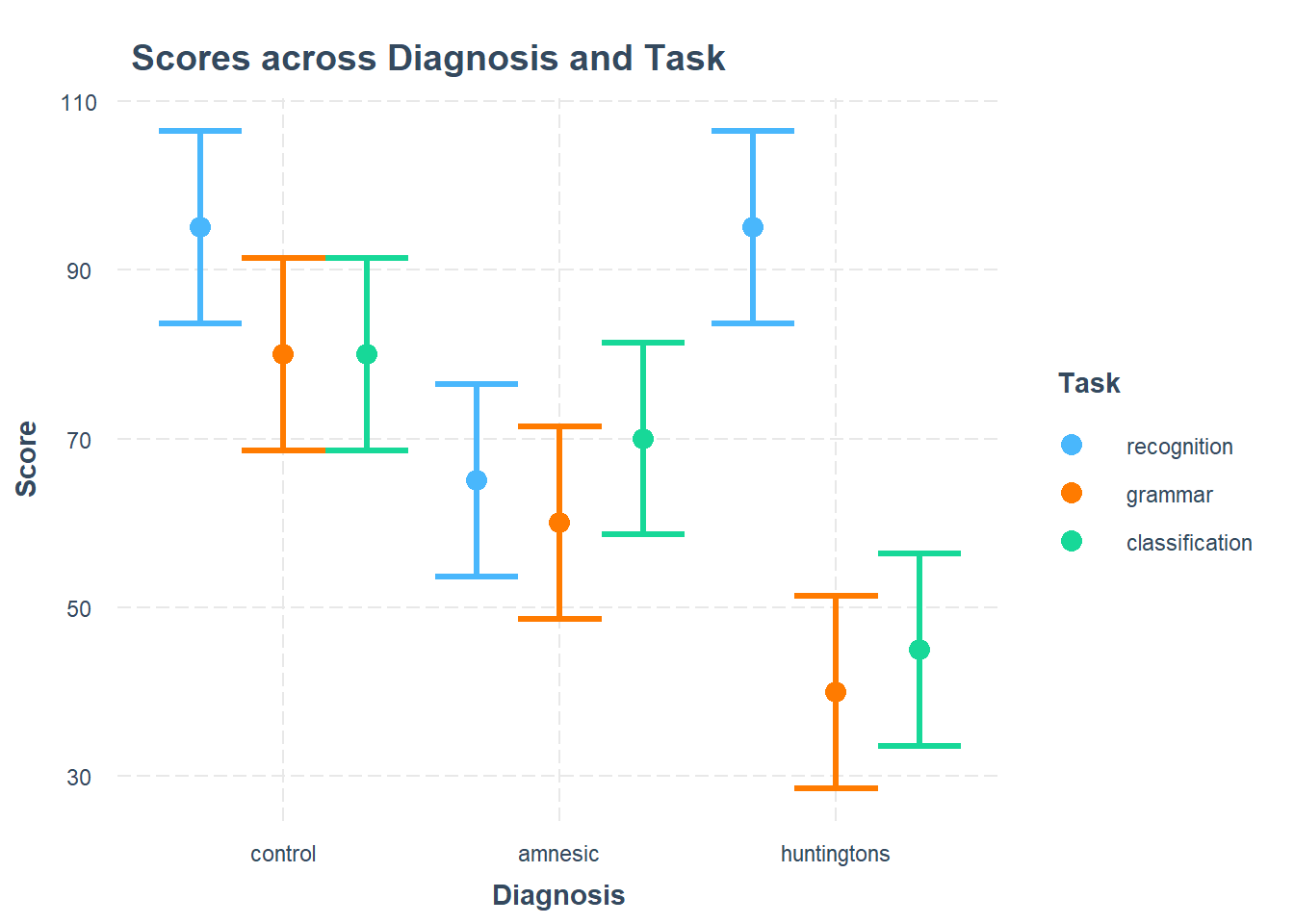
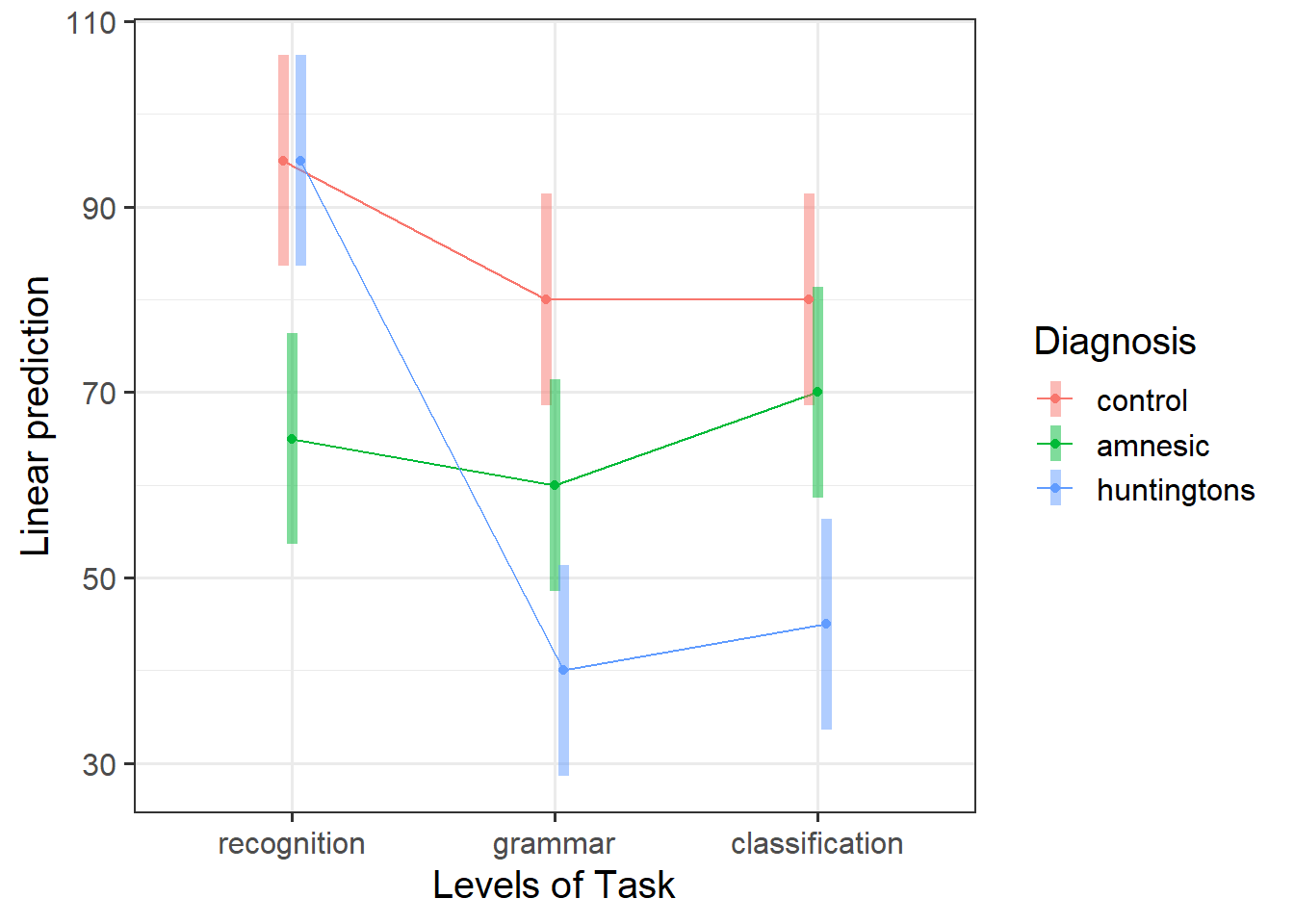
Visualise the interaction, displaying two plots: one with Diagnosis on the x-axis, and the other with Task on the x-axis.
Considering the simple effects that you noted above, identify the significant effects and match them to the corresponding points of your interaction plot.
For an overview and example, review the interaction models > categorical x categorical example > simple effects flashcards.
Recall that the patchwork package allows us to arrange multiple plots using either / or | or +.
Compile Report
Knit your report to PDF, and check over your work. To do so, you should make sure:
- Only the output you want your reader to see is visible (e.g., do you want to hide your code?)
- Check that the tinytex package is installed
- Ensure that the ‘yaml’ (bit at the very top of your document) looks something like this:
---
title: "this is my report title"
author: "B1234506"
date: "07/09/2025"
output: bookdown::pdf_document2
---If you are having issues knitting directly to PDF, try the following:
- Knit to HTML file
- Open your HTML in a web-browser (e.g. Chrome, Firefox)
- Print to PDF (Ctrl+P, then choose to save to PDF)
- Open file to check formatting
To not show the code of an R code chunk, and only show the output, write:
```{r, echo=FALSE}
# code goes here
```To show the code of an R code chunk, but hide the output, write:
```{r, results='hide'}
# code goes here
```To hide both code and output of an R code chunk, write:
```{r, include=FALSE}
# code goes here
```You must make sure you have tinytex installed in R so that you can “Knit” your Rmd document to a PDF file:
install.packages("tinytex")
tinytex::install_tinytex()