Introduction to the Linear Model
DPUK Spring Academy
Department of Psychology
The University of Edinburgh
2025-04-01
Overview
- Day 2: What is a linear model?
- Day 3: But I have more variables, what now?
- Day 4: Interactions
- Day 5: Is my model any good?
Day 3
But I have more variables, now what?
- Part 1 - Quick Recap
- Part 2 - Explaining variance
- Part 3 - More predictors
- Part 4 - More variance explained? (model comparison)
- Part 5 - Coefficients
Quick recap
Models
deterministic
given the same input, deterministic functions return exactly the same output
area of sphere = \(4\pi r^2\)
height of fall = \(1/2 g t^2\)
statistical
\[ \color{red}{\textrm{outcome}} \color{black}{=} \color{blue}{(\textrm{model})} + \color{black}{\textrm{error}} \]
handspan = height + randomness
cognitive test score = age + premorbid IQ + … + randomness
The Linear Model
\[ \begin{align} \color{red}{\textrm{outcome}} & = \color{blue}{(\textrm{model})} + \textrm{error} \\ \qquad \\ \qquad \\ \color{red}{y_i} & = \color{blue}{b_0 \cdot{} 1 + b_1 \cdot{} x_i} + \varepsilon_i \\ \qquad \\ & \text{where } \varepsilon_i \sim N(0, \sigma) \text{ independently} \\ \end{align} \]
The Linear Model
Our proposed model of the world:
\(\color{red}{y_i} = \color{blue}{b_0 \cdot{} 1 + b_1 \cdot{} x_i} + \varepsilon_i\)
The Linear Model
Our model \(\hat{\textrm{f}}\textrm{itted}\) to some data:
\(\hat{y}_i = \color{blue}{\hat b_0 \cdot{} 1 + \hat b_1 \cdot{} x_i}\)
For the \(i^{th}\) observation:
- \(\color{red}{y_i}\) is the value we observe for \(x_i\)
- \(\hat{y}_i\) is the value the model predicts for \(x_i\)
- \(\color{red}{y_i} = \hat{y}_i + \hat\varepsilon_i\)
An example
Our model \(\hat{\textrm{f}}\textrm{itted}\) to some data:
\(\color{red}{y_i} = \color{blue}{5 \cdot{} 1 + 2 \cdot{} x_i} + \hat\varepsilon_i\)
For the observation \(x_i = 1.2, \; y_i = 9.9\):
\[ \begin{align} \color{red}{9.9} & = \color{blue}{5 \cdot{}} 1 + \color{blue}{2 \cdot{}} 1.2 + \hat\varepsilon_i \\ & = 7.4 + \hat\varepsilon_i \\ & = 7.4 + 2.5 \\ \end{align} \]
Categorical Predictors
| y | x |
|---|---|
| 7.99 | Category1 |
| 4.73 | Category0 |
| 3.66 | Category0 |
| 3.41 | Category0 |
| 5.75 | Category1 |
| 5.66 | Category0 |
| ... | ... |
Null Hypothesis Testing
Null Hypothesis Testing
test of individual parameters
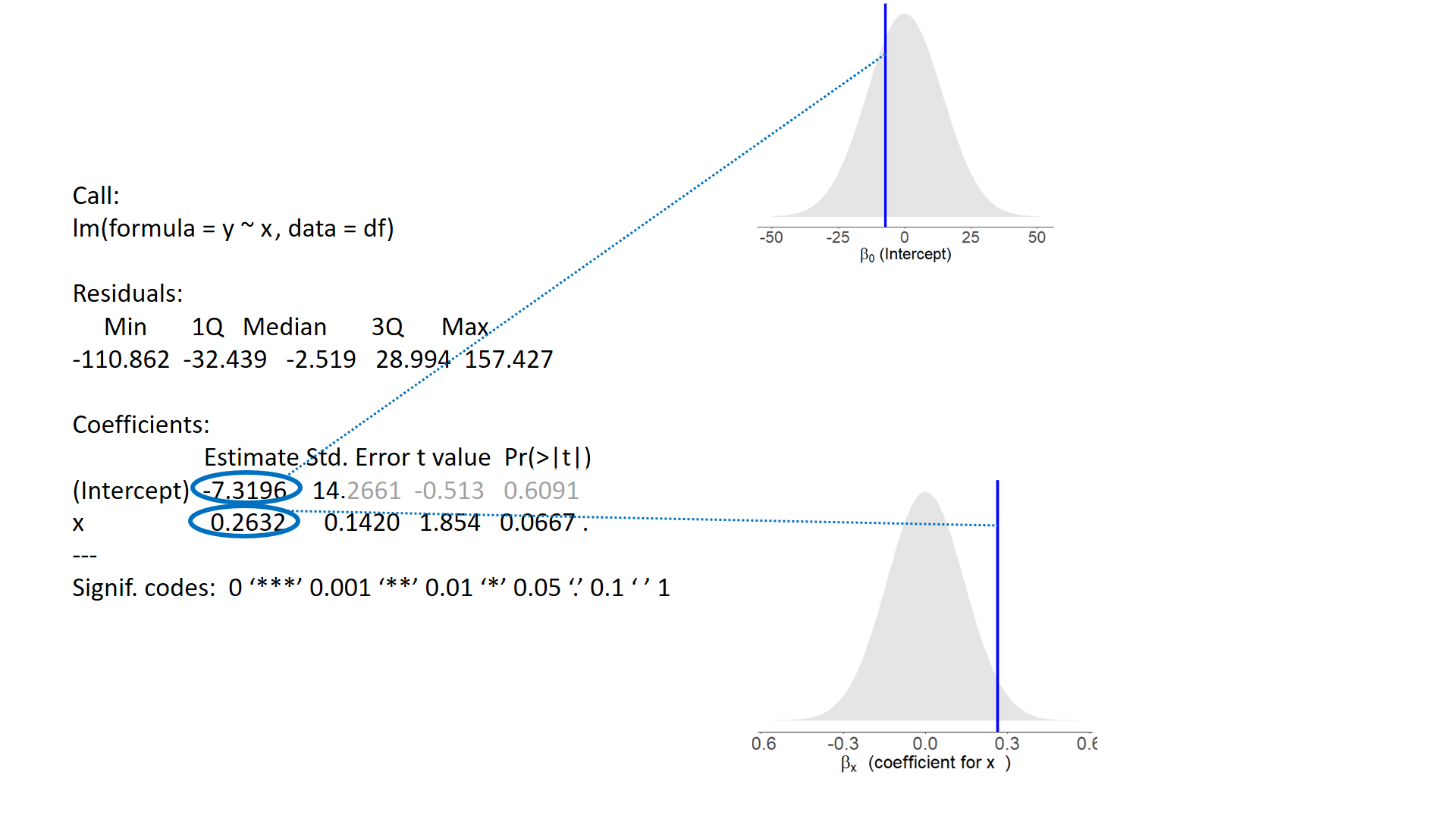
test of individual parameters (2)
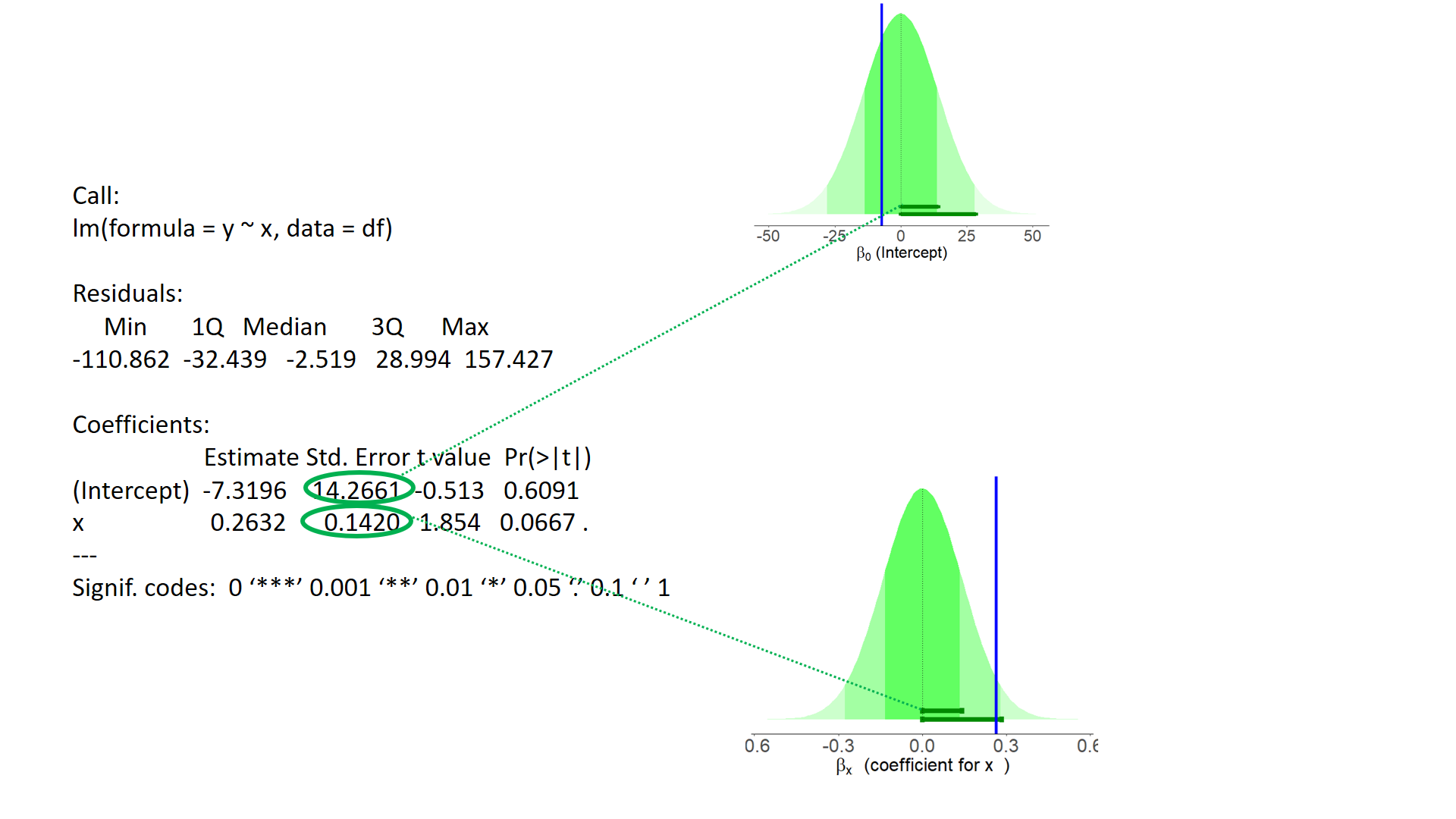
test of individual parameters (3)
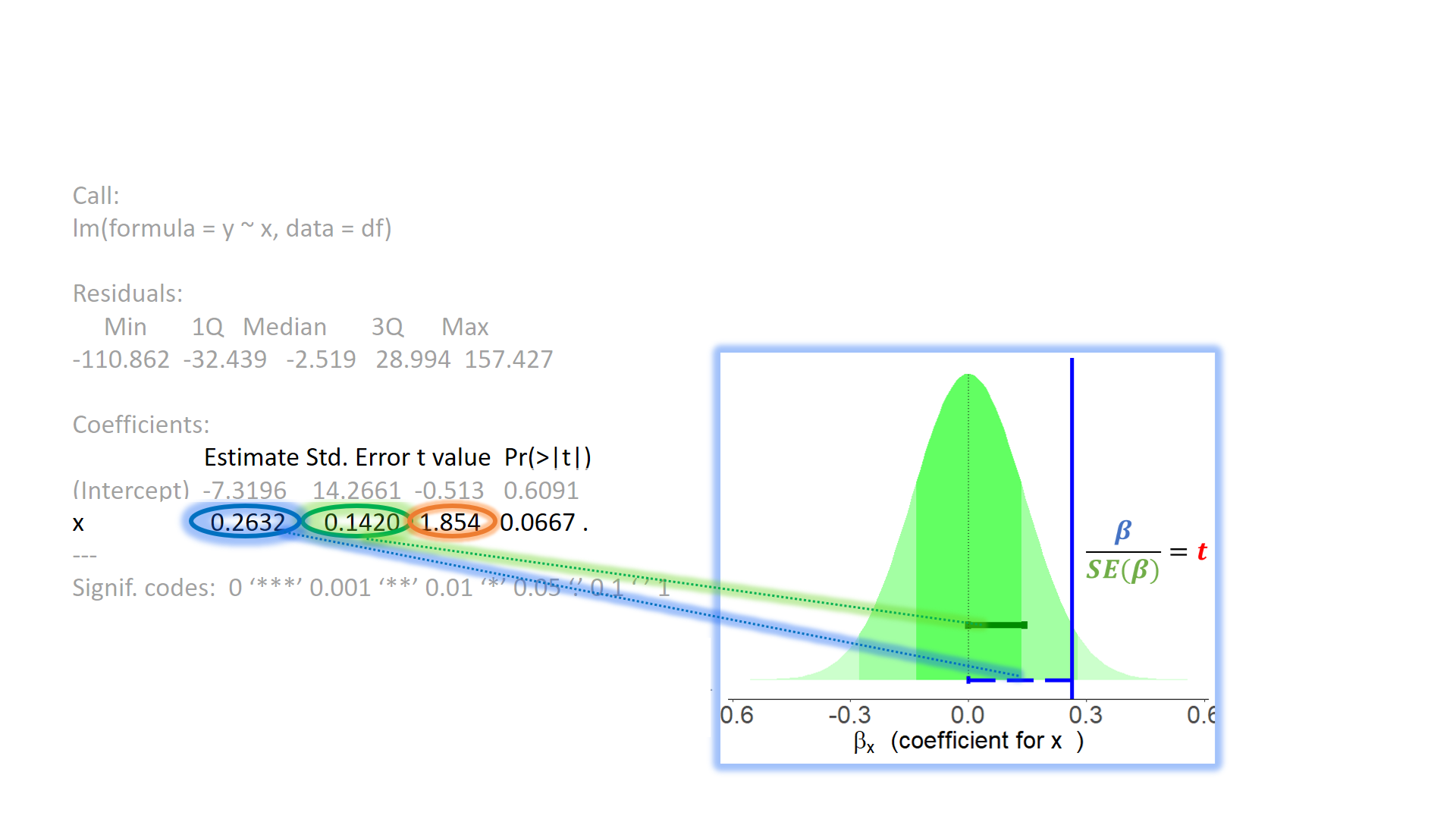
test of individual parameters (4)
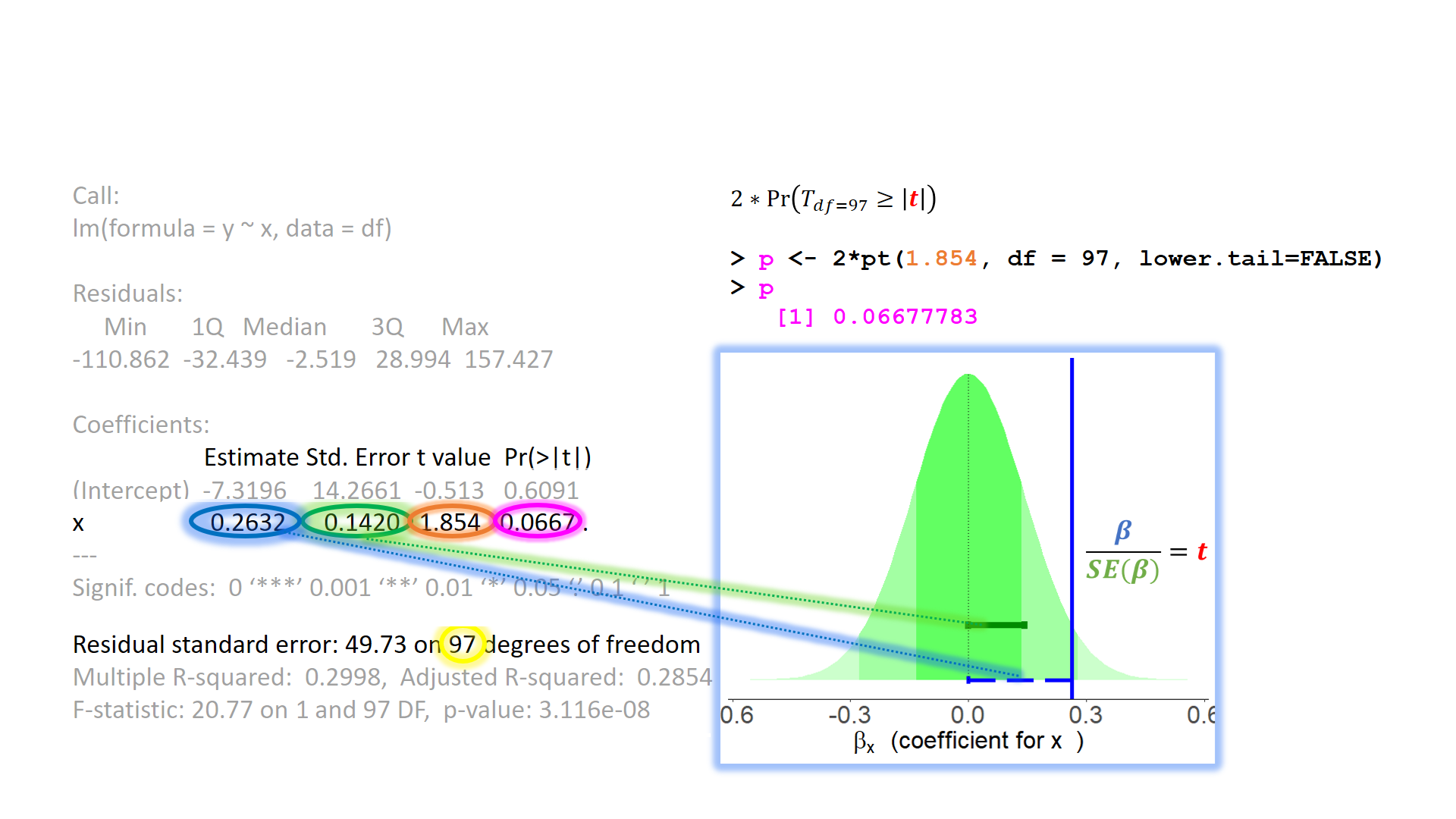
The linear model as variance explained
Sums of Squares
Rather than focussing on slope coefficients, we can also think of our model in terms of sums of squares (SS).
\(SS_{total} = \sum^{n}_{i=1}(y_i - \bar y)^2\)
\(SS_{model} = \sum^{n}_{i=1}(\hat y_i - \bar y)^2\)
\(SS_{residual} = \sum^{n}_{i=1}(y_i - \hat y_i)^2\)
\(R^2\)
F-ratio
\[ \begin{align} & F_{df_{model},df_{residual}} = \frac{MS_{Model}}{MS_{Residual}} = \frac{SS_{Model}/df_{Model}}{SS_{Residual}/df_{Residual}} \\ & \quad \\ & \text{Where:} \\ & df_{model} = k \\ & df_{residual} = n-k-1 \\ & n = \text{sample size} \\ & k = \text{number of explanatory variables} \\ \end{align} \]
...
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) ... ... ... ...
x ... ... ... ...
...
...
Multiple R-squared: 0.66, Adjusted R-squared: 0.657
F-statistic: 190 on 1 and 98 DF, p-value: <2e-16\(F\)-ratio tests the null hypothesis that all the regression slopes in a model are zero
a ratio of the explained to unexplained variance
Mean squares are sums of squares calculations divided by the associated degrees of freedom.
The degrees of freedom are defined by the number of “independent” values associated with the different calculations.
Bigger \(F\)-ratios indicate better models.
- It means the model variance is big compared to the residual variance.
F-test
testing the F-ratio
The null hypothesis for the model says that the best guess of any individuals \(y\) value is the mean of \(y\) plus error.
- Or, that the \(x\) variables carry no information collectively about \(y\).
\(F\)-ratio will be close to 1 when the null hypothesis is true
- If there is equivalent residual to model variation, \(F\)=1
- If there is more model than residual \(F\) > 1
- If there is equivalent residual to model variation, \(F\)=1
\(F\)-ratio is then evaluated against an \(F\)-distribution with \(df_{Model}\) and \(df_{Residual}\) and a pre-defined \(\alpha\)
how does this all help?
The aim of a linear model is to explain variance in an outcome.
In simple linear models, we have a single predictor, but the model can accommodate (in principle) any number of predictors.
However, when we include multiple predictors, those predictors are likely to correlate
Thus, a linear model with multiple predictors finds the optimal prediction of the outcome from several predictors, taking into account their redundancy with one another
how does this all help?
how does \(R^2\) change with the addition of our predictor of interest beyond the variance explained by other variables
does the inclusion of our predictor of interest significantly improve model fit beyond the variance explained by other variables
More predictors
Uses of multiple regression
For prediction: multiple predictors may lead to improved prediction.
For theory testing: often our theories suggest that multiple variables together contribute to variation in an outcome
For covariate control: we might want to assess the effect of a specific predictor, controlling for the influence of others.
- E.g., effects of personality on health after removing the effects of age and sex
![]() Third variables
Third variables
![]() Third variables
Third variables
- X and Y are ‘orthogonal’ (perfectly uncorrelated)
![]() Third variables
Third variables
- X and Y are correlated.
- a = portion of Y’s variance shared with X
- e = portion of Y’s variance unrelated to X
![]() Third variables
Third variables
- X and Y are correlated.
- a = portion of Y’s variance shared with X
- e = portion of Y’s variance unrelated to X
- Z is also related to Y (c)
- Z is orthogonal to X (no overlap)
- relation between X and Y is unaffected (a)
- unexplained variance in Y (e) is reduced, so a:e ratio is greater.
Design is so important! If possible, we could design it so that X and Z are orthogonal (in the long run) by e.g., randomisation.
![]() Third variables
Third variables
- X and Y are correlated.
- Z is also related to Y (c + b)
- Z is related to X (b + d)
Association between X and Y is changed if we adjust for Z (a is smaller than previous slide), because there is a bit (b) that could be attributed to Z instead.
- multiple regression coefficients for X and Z are like areas a and c (scaled to be in terms of ‘per unit change in the predictor’)
- total variance explained by both X and Z is a+b+c
I have control issues..
..and so should you
what do we mean by “control”?
- often quite a vague/nebulous idea
relationship between x and y …
- controlling for z
- accounting for z
- conditional upon z
- holding constant z
- adjusting for z
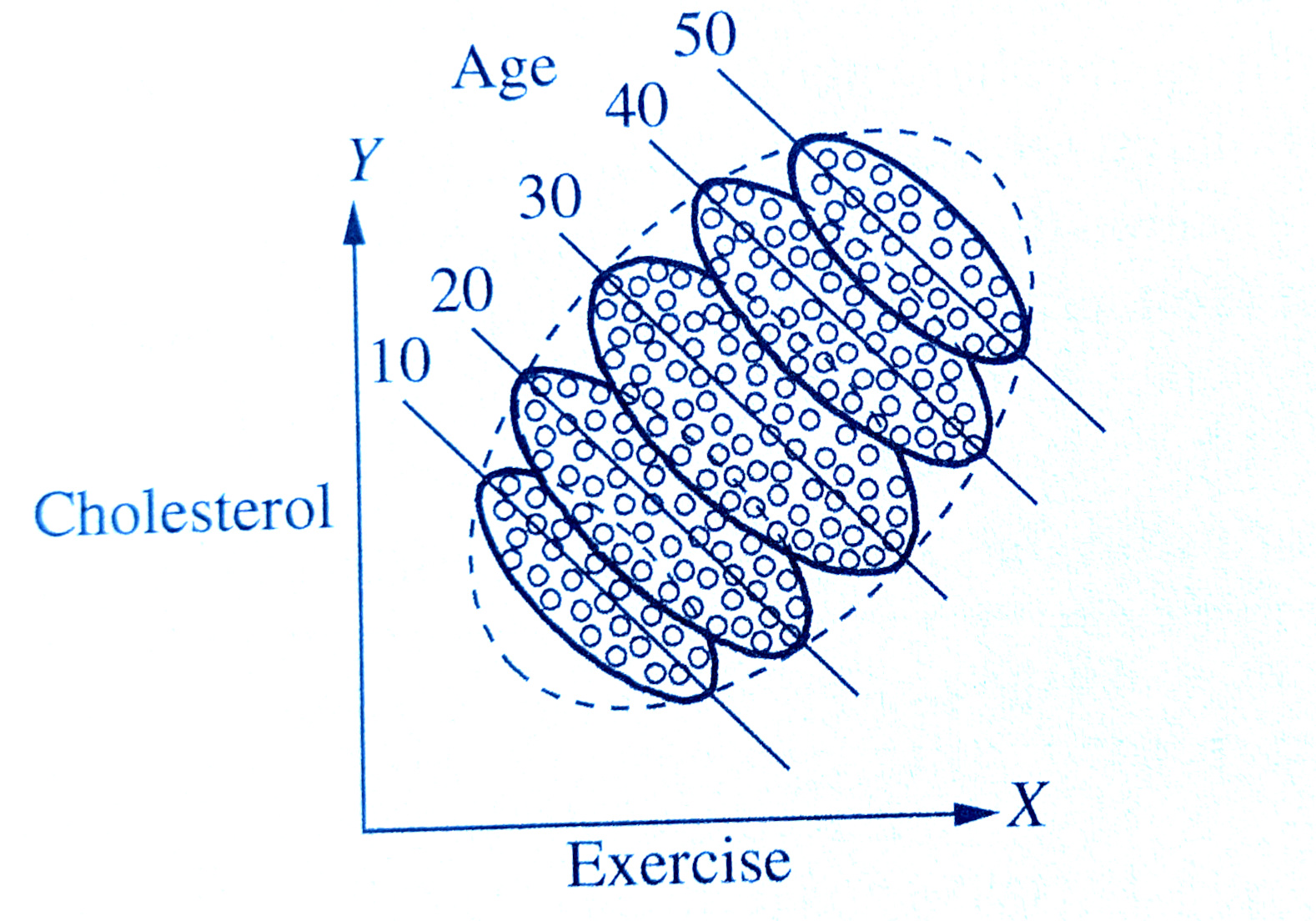
I have control issues..
..and so should you
linear regression models provide “linear adjustment” of Z
sort of like stratification across groups, but Z can now be continuous
assumes effect of X on Y is constant across Z
- (but doesn’t have to, as we’ll see tomorrow)
I have control issues..
..and so should you
In order to estimate “the effect” of X on Y, deciding on what to control for MUST rely on a theoretical model of causes - i.e. a theory about the underlying data generating process
thus far - lm(y~x)
what do i do?
Z is a confounder
Z is a mediator
Z is a collider
example
example
example
example
example 2
- to play around with this, see https://colliderbias.herokuapp.com/
example 2
- to play around with this, see https://colliderbias.herokuapp.com/
example 2
- to play around with this, see https://colliderbias.herokuapp.com/
general heuristics (NOT rules)
control for a confounder
don’t control for a mediator
don’t control for a collider
draw your theoretical model before you get your data, so that you know what variables you (ideally) need.
More variance explained?
More than one predictor?
Model comparison
does
x2explain more variability inythan we would expect by a random variable, over and above variability explained byx1?- once we know x1, does knowing about x2 tell us anything new about y?
tests of multiple parameters
Model comparisons:
tests of multiple parameters (2)
isolate the improvement in model fit due to inclusion of additional parameters
tests of multiple parameters (3)
Test everything in the model all at once by comparing it to a ‘null model’ with no predictors:
Coefficients
multiple regression model
multiple regression coefficients
Call:
lm(formula = y ~ x1 + x2, data = mydata)
Residuals:
Min 1Q Median 3Q Max
-28.85 -6.27 -0.19 6.97 26.05
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 20.372 1.124 18.12 <2e-16 ***
x1 1.884 1.295 1.46 0.1488
x2 2.042 0.624 3.28 0.0015 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 11.2 on 97 degrees of freedom
Multiple R-squared: 0.135, Adjusted R-squared: 0.118
F-statistic: 7.6 on 2 and 97 DF, p-value: 0.00086| term | est | interpretation |
|---|---|---|
| (Intercept) | 20.37 | estimated y when all predictors are zero/reference level |
| x1 | 1.88 | estimated change in y when x1 increases by 1, and all other predictors are held constant |
| x2 | 2.04 | estimated change in y when x2 increases by 1, and all other predictors are held constant |
change in Y for a 1 unit change in X
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -3.8755 2.3911 -1.62 0.1437
age 0.2187 0.0633 3.46 0.0086 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 3.08 on 8 degrees of freedom
Multiple R-squared: 0.599, Adjusted R-squared: 0.549
F-statistic: 11.9 on 1 and 8 DF, p-value: 0.00862unit level predictions
Data
# A tibble: 10 × 4
age exercise therapy stress
<dbl> <dbl> <dbl> <dbl>
1 18 6 0 0.958
2 19 6 0 0.0447
3 20 4 1 -1.81
4 33 5 0 7.60
5 67 3 1 7.53
6 24 1 1 1.61
7 37 7 0 6.03
8 55 6 0 12.6
9 41 4 1 2.60
10 31 5 1 -0.500 Predictions
# A tibble: 10 × 1
prediction
<dbl>
1 0.0615
2 0.280
3 0.499
4 3.34
5 10.8
6 1.37
7 4.22
8 8.15
9 5.09
10 2.90 unit level counterfactuals
Data
# A tibble: 10 × 4
age exercise therapy stress
<dbl> <dbl> <dbl> <dbl>
1 18 6 0 0.958
2 19 6 0 0.0447
3 20 4 1 -1.81
4 33 5 0 7.60
5 67 3 1 7.53
6 24 1 1 1.61
7 37 7 0 6.03
8 55 6 0 12.6
9 41 4 1 2.60
10 31 5 1 -0.500 Predictions
# A tibble: 10 × 1
prediction
<dbl>
1 0.0615
2 0.280
3 0.499
4 3.34
5 10.8
6 1.37
7 4.22
8 8.15
9 5.09
10 2.90 Counterfactuals
# A tibble: 10 × 3
counterfactual_age prediction diff
<dbl> <dbl> <dbl>
1 19 0.280 0.219
2 20 0.499 0.219
3 21 0.718 0.219
4 34 3.56 0.219
5 68 11.0 0.219
6 25 1.59 0.219
7 38 4.44 0.219
8 56 8.37 0.219
9 42 5.31 0.219
10 32 3.12 0.219unit level counterfactuals
Data
# A tibble: 10 × 4
age exercise therapy stress
<dbl> <dbl> <dbl> <dbl>
1 18 6 0 0.958
2 19 6 0 0.0447
3 20 4 1 -1.81
4 33 5 0 7.60
5 67 3 1 7.53
6 24 1 1 1.61
7 37 7 0 6.03
8 55 6 0 12.6
9 41 4 1 2.60
10 31 5 1 -0.500 Predictions
# A tibble: 10 × 1
prediction
<dbl>
1 5.45
2 5.45
3 1.89
4 5.45
5 1.89
6 1.89
7 5.45
8 5.45
9 1.89
10 1.89Counterfactuals
# A tibble: 10 × 3
counterfactual_therapy prediction diff
<dbl> <dbl> <dbl>
1 1 1.89 -3.57
2 1 1.89 -3.57
3 0 5.45 3.57
4 1 1.89 -3.57
5 0 5.45 3.57
6 0 5.45 3.57
7 1 1.89 -3.57
8 1 1.89 -3.57
9 0 5.45 3.57
10 0 5.45 3.57holding constant
(Intercept) age exercise therapy
3.087 0.241 -0.908 -6.940 Data
# A tibble: 10 × 4
age exercise therapy stress
<dbl> <dbl> <dbl> <dbl>
1 18 6 0 0.958
2 19 6 0 0.0447
3 20 4 1 -1.81
4 33 5 0 7.60
5 67 3 1 7.53
6 24 1 1 1.61
7 37 7 0 6.03
8 55 6 0 12.6
9 41 4 1 2.60
10 31 5 1 -0.500 Predictions
# A tibble: 10 × 1
prediction
<dbl>
1 1.98
2 2.22
3 -2.66
4 6.51
5 9.58
6 1.03
7 5.65
8 10.9
9 2.40
10 -0.916Counterfactuals
# A tibble: 10 × 5
age exercise counterfactual_therapy prediction diff
<dbl> <dbl> <dbl> <dbl> <dbl>
1 18 6 1 -4.96 -6.94
2 19 6 1 -4.72 -6.94
3 20 4 0 4.28 6.94
4 33 5 1 -0.434 -6.94
5 67 3 0 16.5 6.94
6 24 1 0 7.97 6.94
7 37 7 1 -1.29 -6.94
8 55 6 1 3.96 -6.94
9 41 4 0 9.34 6.94
10 31 5 0 6.02 6.94Think in hypotheticals
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -3.8755 2.3911 -1.62 0.1437
age 0.2187 0.0633 3.46 0.0086 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 3.08 on 8 degrees of freedom
Multiple R-squared: 0.599, Adjusted R-squared: 0.549
F-statistic: 11.9 on 1 and 8 DF, p-value: 0.00862Think in hypotheticals
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -3.8755 2.3911 -1.62 0.1437
age 0.2187 0.0633 3.46 0.0086 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 3.08 on 8 degrees of freedom
Multiple R-squared: 0.599, Adjusted R-squared: 0.549
F-statistic: 11.9 on 1 and 8 DF, p-value: 0.00862Think in hypotheticals
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -3.8755 2.3911 -1.62 0.1437
age 0.2187 0.0633 3.46 0.0086 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 3.08 on 8 degrees of freedom
Multiple R-squared: 0.599, Adjusted R-squared: 0.549
F-statistic: 11.9 on 1 and 8 DF, p-value: 0.00862Think in hypotheticals
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 5.45 1.99 2.75 0.025 *
therapy -3.57 2.81 -1.27 0.240
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 4.44 on 8 degrees of freedom
Multiple R-squared: 0.168, Adjusted R-squared: 0.0637
F-statistic: 1.61 on 1 and 8 DF, p-value: 0.24Think in hypotheticals
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 3.0874 3.1338 0.99 0.36259
age 0.2412 0.0334 7.22 0.00036 ***
exercise -0.9082 0.4817 -1.89 0.10836
therapy -6.9398 1.6246 -4.27 0.00525 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1.61 on 6 degrees of freedom
Multiple R-squared: 0.918, Adjusted R-squared: 0.877
F-statistic: 22.3 on 3 and 6 DF, p-value: 0.00118Think in hypotheticals
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 3.0874 3.1338 0.99 0.36259
age 0.2412 0.0334 7.22 0.00036 ***
exercise -0.9082 0.4817 -1.89 0.10836
therapy -6.9398 1.6246 -4.27 0.00525 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1.61 on 6 degrees of freedom
Multiple R-squared: 0.918, Adjusted R-squared: 0.877
F-statistic: 22.3 on 3 and 6 DF, p-value: 0.00118Think in hypotheticals
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 3.0874 3.1338 0.99 0.36259
age 0.2412 0.0334 7.22 0.00036 ***
exercise -0.9082 0.4817 -1.89 0.10836
therapy -6.9398 1.6246 -4.27 0.00525 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1.61 on 6 degrees of freedom
Multiple R-squared: 0.918, Adjusted R-squared: 0.877
F-statistic: 22.3 on 3 and 6 DF, p-value: 0.00118categorical predictors
example - simple regression
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.6027 0.0347 17.35 < 2e-16 ***
isMonkeyYES -0.2599 0.0449 -5.79 0.0000018 ***(Intercept): the estimated brain mass of Humans (when the isMonkeyYES variable is zero)isMonkeyYES: the estimated change in brain mass from Humans to Monkeys (change in brain mass when moving from isMonkeyYES = 0 to isMonkeyYES = 1).
example - simple regression
example - in multiple regression
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.66831 0.05048 13.24 1.6e-14 ***
age -0.00408 0.00234 -1.75 0.09 .
isMonkeyYES -0.24693 0.04415 -5.59 3.5e-06 ***(Intercept): the estimated brain mass of new-born Humans (when both age is zero and isMonkeyYES is zero)age: the estimated change in brain mass for every 1 year increase in age, holding isMonkey constant.isMonkeyYES: the estimated change in brain mass from Humans to Monkeys (change in brain mass when moving from isMonkeyYES = 0 to isMonkeyYES = 1), holding age constant.
example - in multiple regression
example - multiple levels
braindata <- braindata %>% mutate(
speciesPotar = ifelse(species == "Potar monkey", 1, 0),
speciesRhesus = ifelse(species == "Rhesus monkey", 1, 0),
)
braindata %>%
select(mass_brain, species, speciesPotar, speciesRhesus) %>%
head()# A tibble: 6 × 4
mass_brain species speciesPotar speciesRhesus
<dbl> <chr> <dbl> <dbl>
1 0.449 Rhesus monkey 0 1
2 0.577 Human 0 0
3 0.349 Potar monkey 1 0
4 0.626 Human 0 0
5 0.316 Potar monkey 1 0
6 0.398 Rhesus monkey 0 1- For a human, both
speciesPotar == 0andspeciesRhesus == 0 - For a Potar monkey,
speciesPotar == 1andspeciesRhesus == 0 - For a Rhesus monkey,
speciesPotar == 0andspeciesRhesus == 1
example - multiple levels
Call:
lm(formula = mass_brain ~ species, data = braindata)
Residuals:
Min 1Q Median 3Q Max
-0.30010 -0.06254 0.00129 0.04779 0.18664
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.6027 0.0275 21.94 < 2e-16 ***
speciesPotar monkey -0.3574 0.0414 -8.63 7.4e-10 ***
speciesRhesus monkey -0.1526 0.0426 -3.59 0.0011 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.103 on 32 degrees of freedom
Multiple R-squared: 0.699, Adjusted R-squared: 0.681
F-statistic: 37.2 on 2 and 32 DF, p-value: 4.45e-09
Call:
lm(formula = mass_brain ~ speciesPotar + speciesRhesus, data = braindata)
Residuals:
Min 1Q Median 3Q Max
-0.30010 -0.06254 0.00129 0.04779 0.18664
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.6027 0.0275 21.94 < 2e-16 ***
speciesPotar -0.3574 0.0414 -8.63 7.4e-10 ***
speciesRhesus -0.1526 0.0426 -3.59 0.0011 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.103 on 32 degrees of freedom
Multiple R-squared: 0.699, Adjusted R-squared: 0.681
F-statistic: 37.2 on 2 and 32 DF, p-value: 4.45e-09example - multiple levels
more levels = more dimensions
example - multiple levels
Call:
lm(formula = mass_brain ~ species, data = braindata)
Residuals:
Min 1Q Median 3Q Max
-0.30010 -0.06254 0.00129 0.04779 0.18664
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.6027 0.0275 21.94 < 2e-16 ***
speciesPotar monkey -0.3574 0.0414 -8.63 7.4e-10 ***
speciesRhesus monkey -0.1526 0.0426 -3.59 0.0011 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.103 on 32 degrees of freedom
Multiple R-squared: 0.699, Adjusted R-squared: 0.681
F-statistic: 37.2 on 2 and 32 DF, p-value: 4.45e-09